I am a mycologist with a passion for understanding how fungal symbioses are established, maintained, and evolve over time. I am a Professor in the Botany and Plant Pathology Department at Oregon State University in Corvallis, OR, USA.
I am a mycologist with a passion for understanding how fungal symbioses are established, maintained, and evolve over time. I am a Professor in the Botany and Plant Pathology Department at Oregon State University in Corvallis, OR, USA.
I am fascinated by the questions: how do fungi interact with the world, and how can they help us?
Current Projects:
-
Research on Fungal diversity and evolution
- Director of my research lab, see Uehlinglab.com.
- OSC Fungal Herbarium Curator
- Mycologist for the Psilocybin Advisory Board
-
Instructor for Mycology (BOT 461/561) and Population Genomics (BDS 477/577)
Research interests
My research offers insight into how fungi function in symbioses with bacteria, plants, and humans and how fungal genomes evolve as a consequence. My research has been supported by The National Science Foundation, The Department of Energy, NASA, and various Mycological Societies. I am currently conducting research in the following areas:
Media features
ESPN E60 documentary ‘Peace of Mind’
National Geographic A psychedelic surprise may be thriving in your local garden
Mushroom Revival Podcast How Fungi & Bacteria Interact
Scientific American Restrictions on Psilocybin ‘Magic Mushrooms’ Are Easing as Research Ramps Up
Science Daily Research helps provide scientific framework for psilocybin use in therapeutic settings
Polk County Observer Oregon’s psilocybin program stands on thousands of years of indigenous experience
Oregon State University News Room Oregon State research helps provide scientific framework for psilocybin use in therapeutic settings
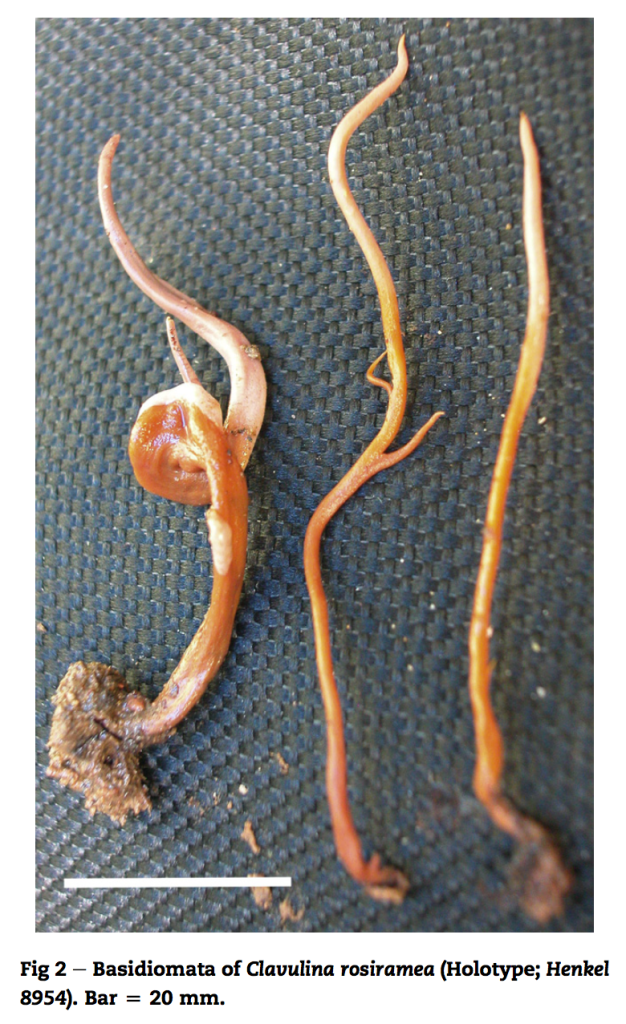
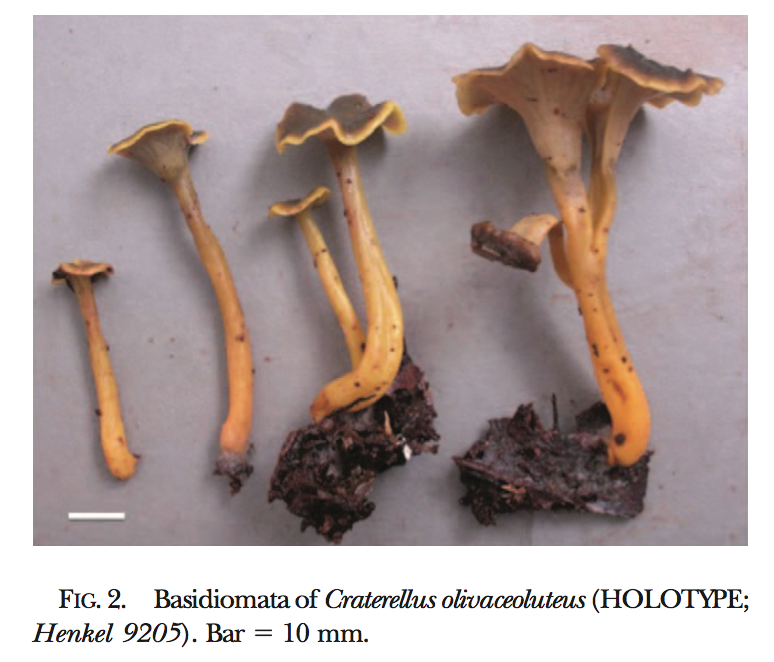
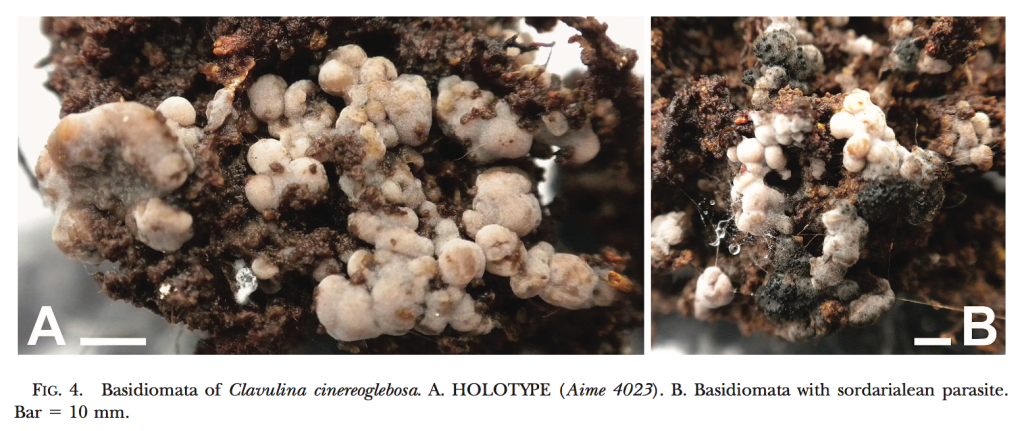
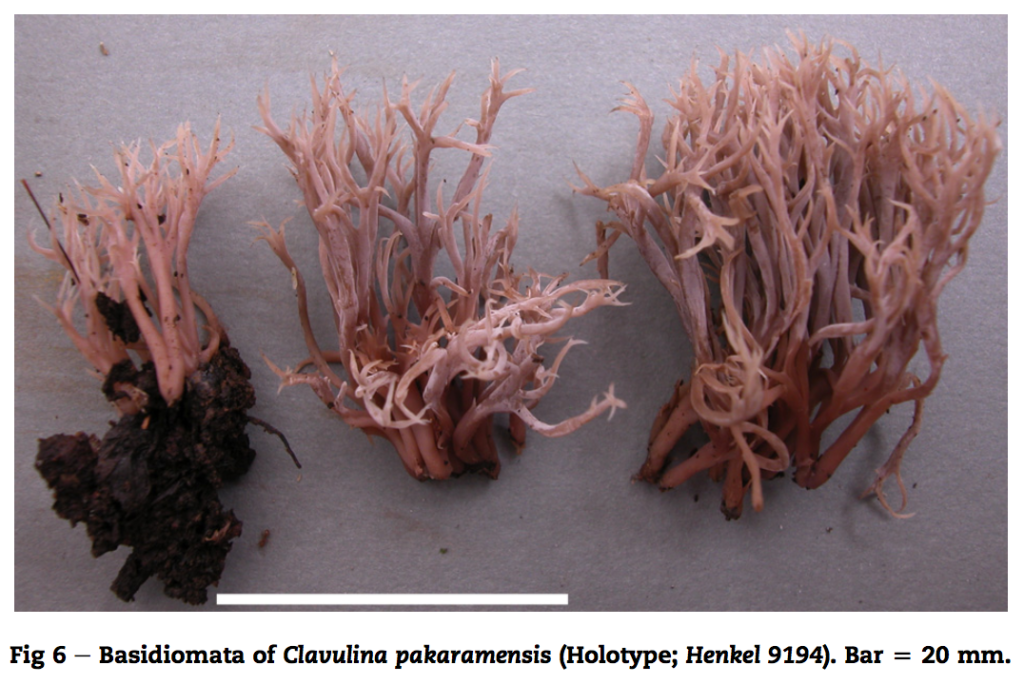
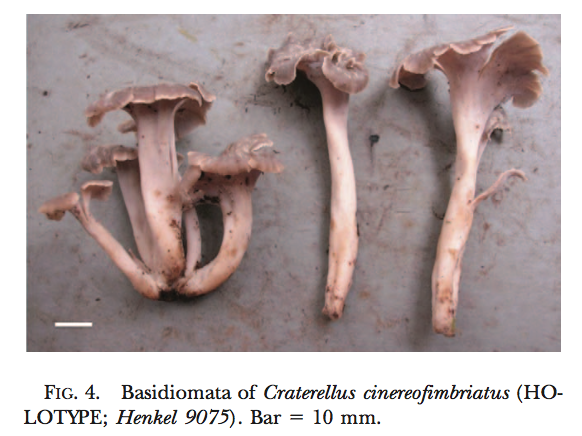
New fungal species descriptions
Below are images of fungal species I have found and described. For more info click here and for associated publications see my CV